Many restriction enzymes are sensitive to the DNA methylation states. Cleavage can be blocked or impaired when a particular base in the recognition site is modified.
Scientists at NEB recently identified the MspJI family of restriction enzymes, which are dependent on methylation and hydroxymethylation for cleavage to occur. These enzymes excise ~ 32 base pair fragments containing a centrally located 5-hmC or 5-mC modified residue that can be extracted and sequenced. Due to the known position of this epigenetic modification, bisulfite conversion is not required prior to downstream analysis.
These EpiMark validated, methylation-dependent restriction enzymes expand the potential for mapping epigenetic modifications and simplify the study of DNA methylation. Additionally, they provide an opportunity to better understand the role of 5-hydroxymethylcytosine in the genome.
A variety of our existing restriction enzymes can also be used to study epigenetic modifications of DNA, such as, DpnI and DpnII that recognize the same sequence, but different methylation patterns. McrBC also only cleaves DNA that is methylated on one or both strands.
- Specificity to epigenetically-relevant DNA modifications (5-mC and 5-hmC)
- Easy-to-use protocols (enzymatic digestion followed by gel extraction)<
- Less harsh than bisulfite conversion
- Simplified data analysis
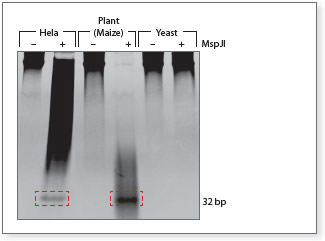
Simplify DNA methylation analysis with MspJI
MspJI recognizes methylated and hydroxymethylated DNA and cleaves out ~ 32 bp fragments for downstream sequencing analysis. Overnight digestion of 1 µg of genomic DNA from various sources with or without MspJI is shown. Note: Yeast DNA does not contain methylated DNA and thus, no 32-mer is detected.
Product table
As of: 01.01.2024
Further information can be found in our Technical Resources section or at neb.com. Information on trademarks can be found here.