EnGen SpRY Cas9
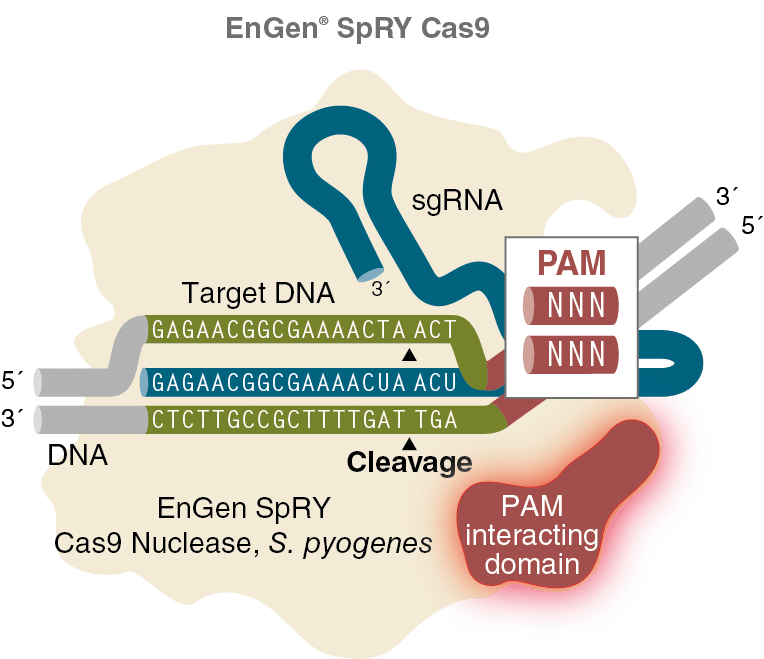
EnGen SpRY Cas9 is a variant of Cas9 nuclease from S. pyogenes with several point mutations within the PAM interacting domain. Unlike wildtype Cas9, EnGen SpRY Cas9 is not constrained by the presence of an NGG PAM and can produce a double stranded break three nucleotides upstream of any trinucleotide sequence in in vitro applications.
EnGen Seq1 Cas9
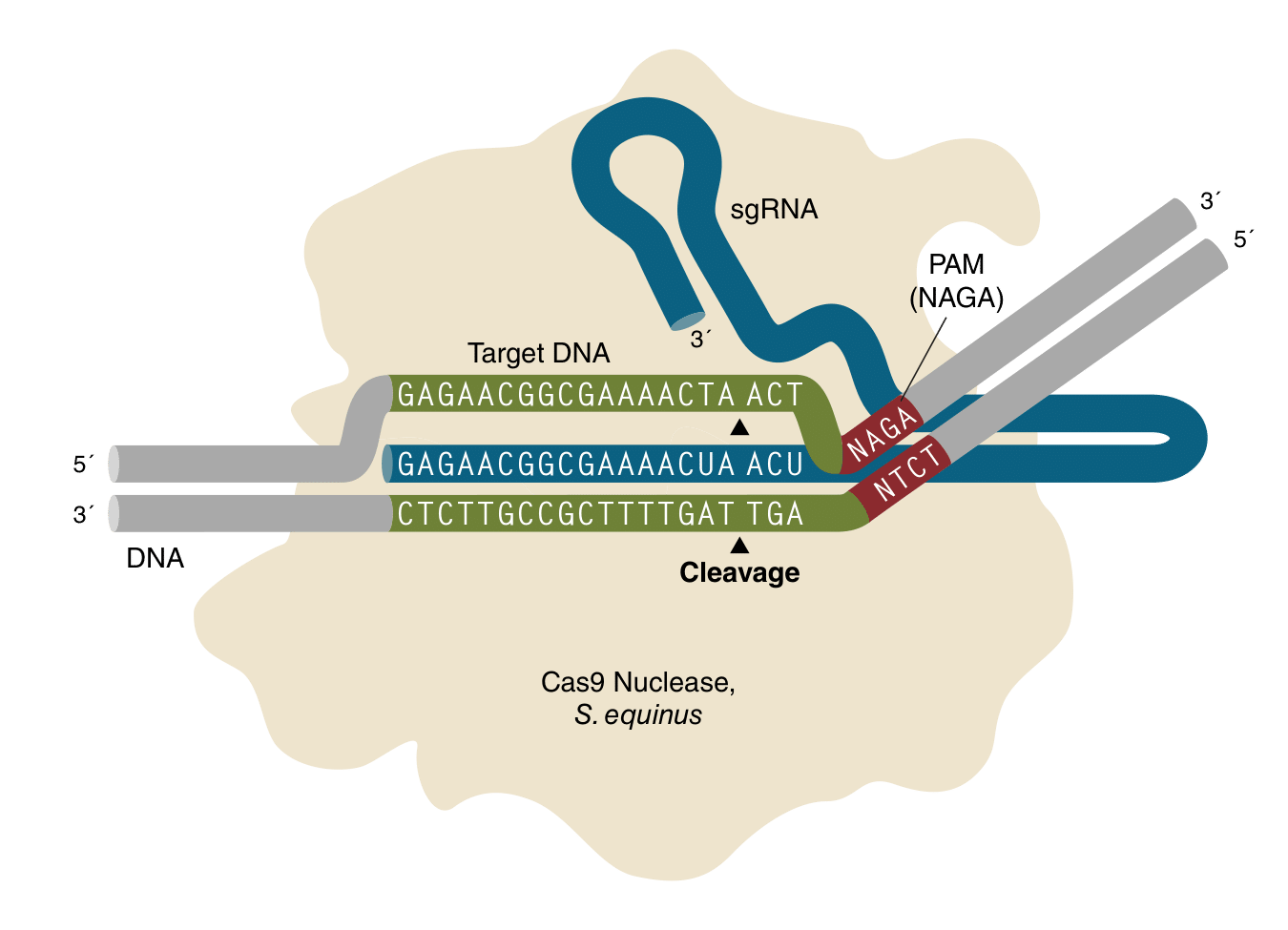
EnGen Seq1 Cas9 forms a ribonucleoprotein complex with a 116-nt single guide RNA (sgRNA), which includes a 20-nt sequence complementary to the target DNA immediately upstream of a 5′–NAGA–3′ protospacer adjacent motif (PAM). DNA cleavage by EnGen Seq1 Cas9 produces a double-stranded break occurring 3 nucleotides upstream of the PAM. EnGen Seq1 Cas9 has Simian virus 40 (SV40) T antigen nuclear localization signals (NLS) at both the N- and C-termini of the protein.
EnGen Spy Cas9 HF1
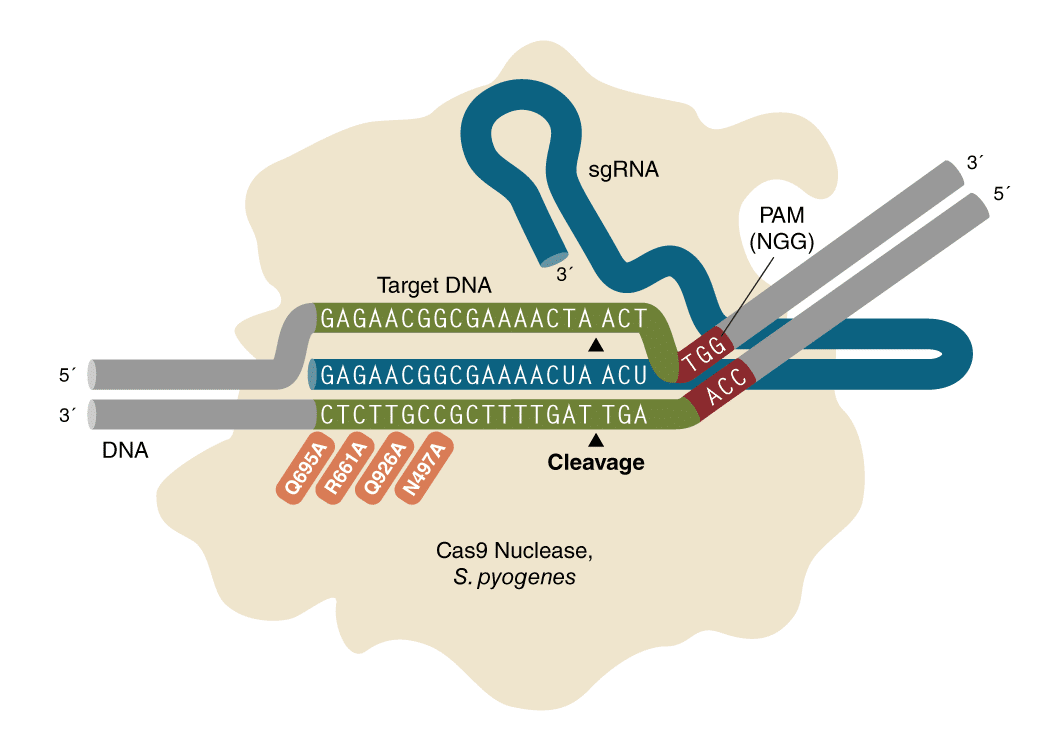
The all-new EnGen Spy Cas9 HF1 is a high-fidelity variant of EnGen Spy Cas9 NLS from Streptococcus pyogenes with reduced non-specific DNA cleavage. The improvement of fidelity was achieved by 4 mutations in the DNA-binding domain. Find out more about this improvement in this Nature publication. EnGen Spy Cas9 HF1 contains two NLS on the N- and C-termini of the protein for high targeting efficiencies.
EnGen Spy Cas9 NLS
EnGen Spy Cas9 NLS, is an RNA-guided endonuclease that catalyzes site- specific cleavage of double stranded DNA. The location of the break is within the target sequence 3 bases from the NGG PAM (Protospacer Adjacent Motif). The PAM sequence, NGG, must follow the targeted region on the opposite strand of the DNA with respect to the region complementary sgRNA sequence. EnGen Spy Cas9 NLS contains Simian virus 40 (SV40) T antigen nuclear localization sequence (NLS) on the N- and C- termini of the protein.
- Deal for direct introduction of Cas9/sgRNA complexes
- Dual NLS for improved transport to the nucleus
- Compatible with EnGen sgRNA Synthesis Kit, S. pyogenes and the EnGen Mutation Detection Kit
For in vitro applications NEB offers recombinant Streptococcus pyogenes Cas9 Nuclease.
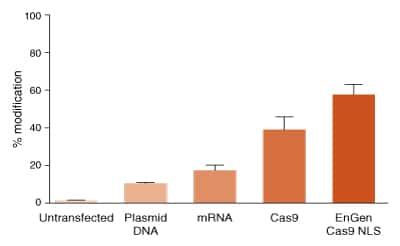
Cas9 and sgRNA targeting a human gene were delivered to HEK293 cells by transfection. Transfected plasmid DNA contained expression cassettes for 2x NLS (N- and C-terminal) Cas9 and sgRNA. Plasmid DNA was delivered using TransIT-X2 (Mirus). Transfected mRNA was modified with pseudouridine and 5-methylcytosine and encoded 2x NLS (Nand C-terminal) Cas9. sgRNA was co-transfected with the mRNA using TransIT-mRNA. Cas9 RNPs were delivered in reverse transfections using Lipofectamine RNAiMAX (Life Technologies) using 10 nanomolar final concentration of ribonucleoprotein (RNP). Cas9 has no NLS in the protein sequence. EnGen Cas9 has N- and C-terminal NLSs. The efficiency of editing was determined using T7 Endonuclease I assay and is expressed as % modification.
NEBs Cas9 in microinjection in customer experiment: in vivo Gene Editing in Zebrafish
|
„The experiments with NEB Cas9 are going well, seems very effective with a variety of sgRNAs. The high conc. Cas9 protein is extremely efficient in vivo!“ James A. Gagnon, Department of Molecular and Cellular Biology, Harvard University, USA |
||||
 |
||||
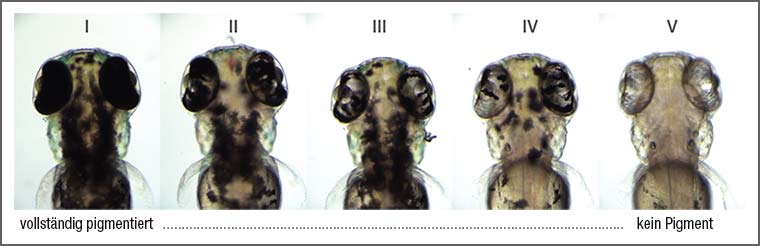
Mutations in the tyrosinase gene block melanocyte pigmentation in zebrafish: 1-cell embryos were injected with 10 sgRNAs targeting tyrosinase and NEB 18 μM Cas9 protein. Categories of pigmentation at 3 days post fertilization. At 2nl injection volume, >90% of embryos completely lack pigmentation.
Images and data by courtesy of James A. Gagnon, Department of Molecular and Cellular Biology, Harvard University, USA
Further references:Nakagawa, et al. Biology Open (2016) 0, 1-7, Ultra-superovulation for the CRISPR-Cas9-mediated production of gene-knockout, single-amino-acid-substituted, and floxed mice
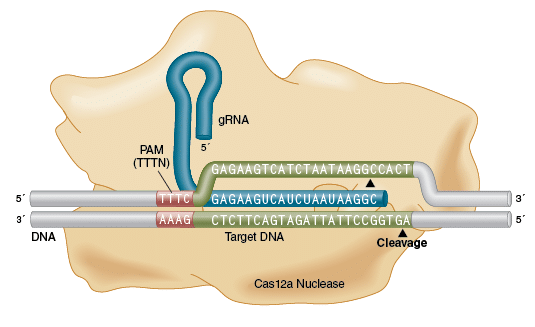
EnGen Lba Cas12a (Cpf1)
EnGen Lba Cas12a (Cpf1) is a programmable DNA endonuclease guided by a single guide RNA (gRNA). Targeting requires a gRNA complementary to the target site as well as a 5′ TTTN protospacer adjacent motif (PAM) on the DNA strand opposite the target sequence. Cleavage by EnGen Lba Cas12a (Cpf1) occurs ~18 bases 3′ of the PAM and leaves 5′ overhanging ends.
- T-rich TTTN PAM sequence opens up additional genomic regions for targeting
- Shorter, 40-44 base guide RNA
- Two nuclear localization signals for improved transport to the nucleus
- 5’ overhanging termini on cleavage products
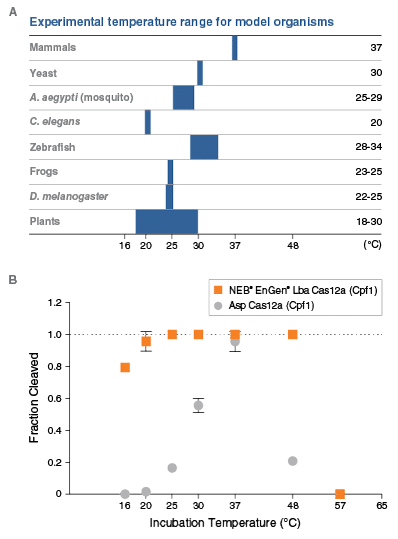
- Active from 16 to 48°C
- Maintains activity at lower temperatures than the Acidaminococcus orthologs, permitting editing in ectothermic organisms such as zebra fish and xenopus
- High concentration liquid format can be used for microinjection, electroporation and lipofection
Purified Lba or Asp Cas12a proteins were used to form RNP complexes with their canonical guide RNAs. The DNA target is a single a PCR product 5´-end labeled with FAM. Lba or Asp Cas12a RNPs were incubated with target DNA at the temperatures indicated at a ratio of 10:1 RNP to DNA for 10 minutes. Cleavage products were resolved by capillary electrophoresis and quantitated by integration of signal corresponding to intact and cleaved fragments. Data are shown as the mean +/- SD of duplicate experiments.
EnGen Spy Cas9 Nickase
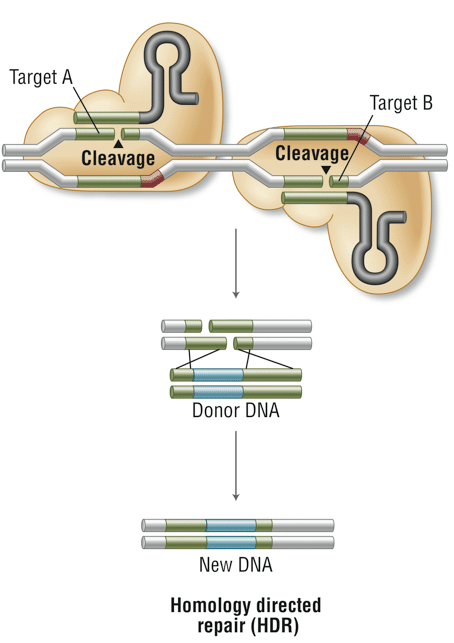
EnGen Spy Cas9 Nickase is a variant of Cas9 nuclease differing by a point mutation (D10A) in the RuvC nuclease domain, which enables it to nick, but not cleave, DNA. Double-stranded DNA breaks can be generated with reduced off-target cleavage by targeting two sites in close proximity (generally 0-20 bp apart) and with PAMs facing outward to leave 5’ overhangs. Nicking by EnGen Spy Cas9 Nickase occurs on the DNA strand opposite the target sequence.
- Variant of Cas9 nuclease differing by a point mutation (D10A) in the RuvC nuclease domain contains SV40 T antigen nuclear localization (NLS) sequences on the N and C- termini.
- Capable of generating nicks, but not cleaving DNA
- DNA double strand breaks can be generated, with reduced off-target cleavage, by targeting two sites with EnGen Cas9 Nickase in close proximity
EnGen Spy dCas9 (SNAP-tag)
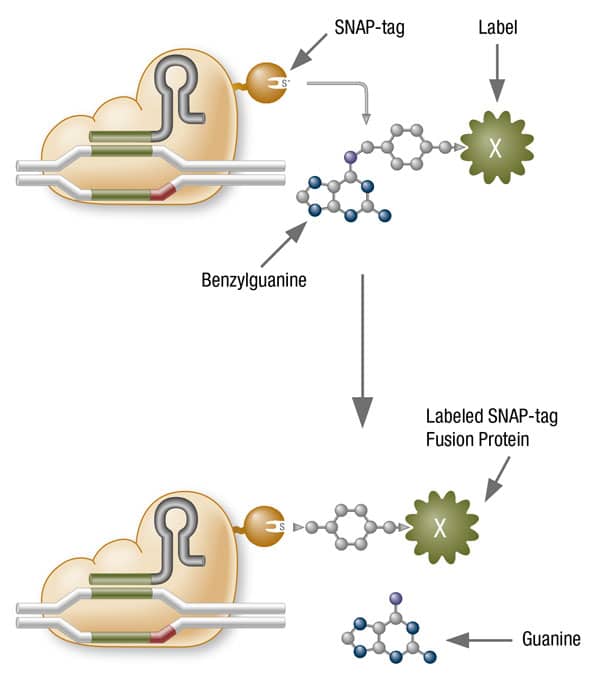
EnGen Spy dCas9 is an inactive mutant of Cas9 nuclease that retains programmable DNA binding activity. EnGen Spy dCas9 contains two point mutations, D10A and H840A, which inactivate its RuvC and HNH nuclease domains, respectively. The N-terminal SNAP-tag allows for covalent attachment of fluorophores, biotin, and a number of other conjugates useful for visualization and target enrichment. Like with Cas9 nuclease, DNA binding by EnGen Spy dCas9 requires a guide RNA complementary to a target sequence with a protospacer adjacent motif (PAM), NGG, downstream of the target sequence.
- An inactive mutant of Cas9 nuclease that retains programmable DNA binding activity
- contains a SV40 T antigen nuclear localization sequence (NLS) at the C- terminus of the protein.
- The N-terminal SNAP-tag allows for covalent attachment of fluorophores, biotin, and a number of other conjugates useful for visualization and target enrichment
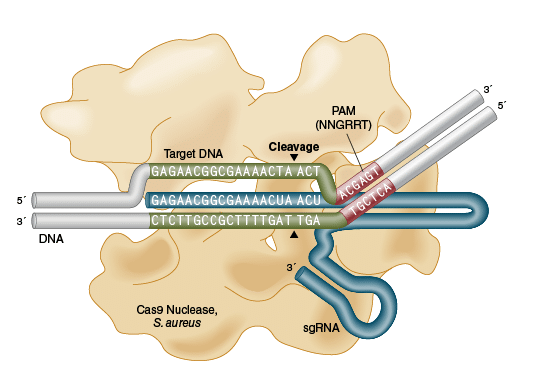
EnGen Sau Cas9
- Cas9 enzyme from Staphylococcus aureus
- RNA-guided DNA endonuclease that catalyzes site-specific cleavage of double-stranded DNA
- Longer 5′ NNGRRT 3′ PAM sequence allows for higher specificity
Abb. rechts: Schematic representation of Sau Cas9 nuclease sequence recognition and DNA cleavage
Available products:
As of: 23.06.2025
Further information can be found in our Technical Resources section or at neb.com. Information on trademarks can be found here.