RNA isolation made easy:
The “universal” Monarch Kit for quick and easy isolation of total RNA from cells, tissue, blood, bacteria, plants, …
The Monarch Spin RNA Isolation Kit (Mini) is a comprehensive solution for sample preservation, cell lysis, gDNA removal, and purification of total RNA from various biological samples, including cultured cells, mammalian tissues, microbes, plants, insects and blood.
With just this single kit, users can purify up to 100 µg of high-quality total RNA from multiple sample types, from standard as well as tough-to-lyse samples, with included enzymes, reagents, and specialized gDNA removal columns. The kit uniquely enables binding capacities like RNA purification miniprep kits, combined with the low elution volumes of micro kits. Cleanup of enzymatic reactions and purification of RNA from TRIzol -extracted samples is also possible using this kit.
- Purify up to 100 µg of high-quality total RNA from multiple sample types including tough-to-lyse samples with this single all-in-one kit
- Extract RNA from samples such as cells, tissues including fibrous or lipid-rich, Gram-positive or Gram-negative bacteria, yeast, plants, insects and more
- Kit includes Proteinase K for sample processing and Monarch StabiLyse™ DNA/RNA Buffer for sample preservation
- Achieve binding capacity of a standard mini kit (100 µg) with the low elution volume of a micro kit (10 µl)
- Remove genomic DNA contamination with specialized columns and RNase-free DNase included in the kit
- Capture total RNA pool in your sample including small RNAs <200 nt
Purified RNA has metrics with A260/280 and A260/230 ratios typically > 2.0, high RNA integrity scores, and minimal residual gDNA. This kit captures RNA ranging in size from full-length rRNAs down to intact miRNAs. Purified RNA is suitable for downstream applications such as RT-qPCR, cDNA synthesis, RNA-seq, and RNA hybridization-based technologies.
Designed with sustainability in mind
The kit offers significantly less plastic through uniquely designed columns and bottles with plastic reduction while still maintaining structural integrity for centrifugation or vacuum manifolds and other steps.
Kit boxes, inserts, and protocol cards are made of recycled paper.
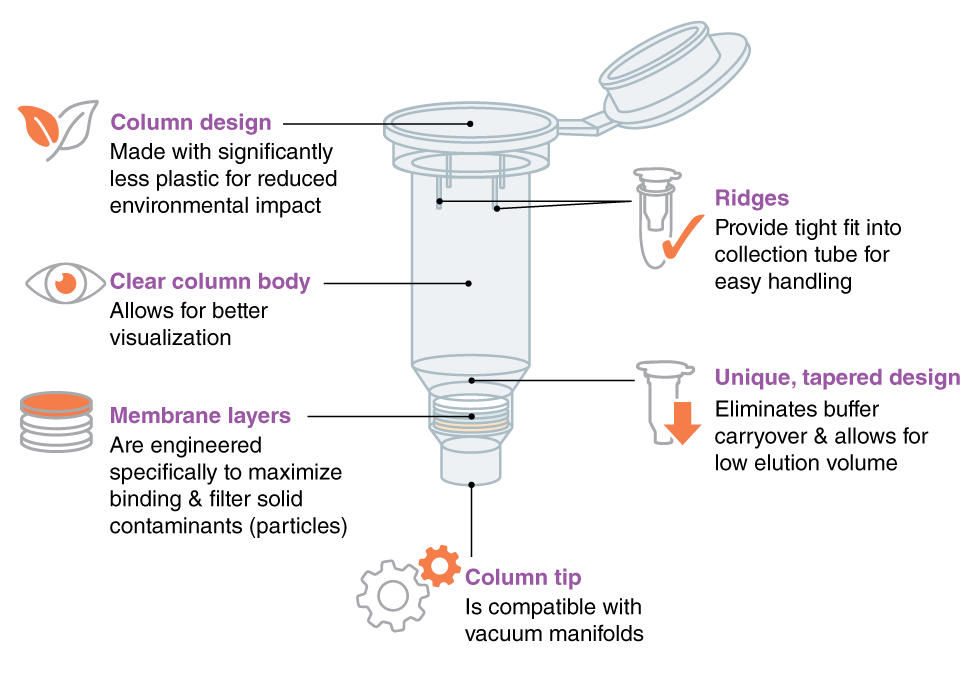
Unique Column Design
NEB Monarch’s unique column design and membrane assembly allows high-quality, highly-concentrated nucleic acid purification with low elution volume, for downstream applications. The column is designed and made with significantly less plastic for a reduced environmental impact.
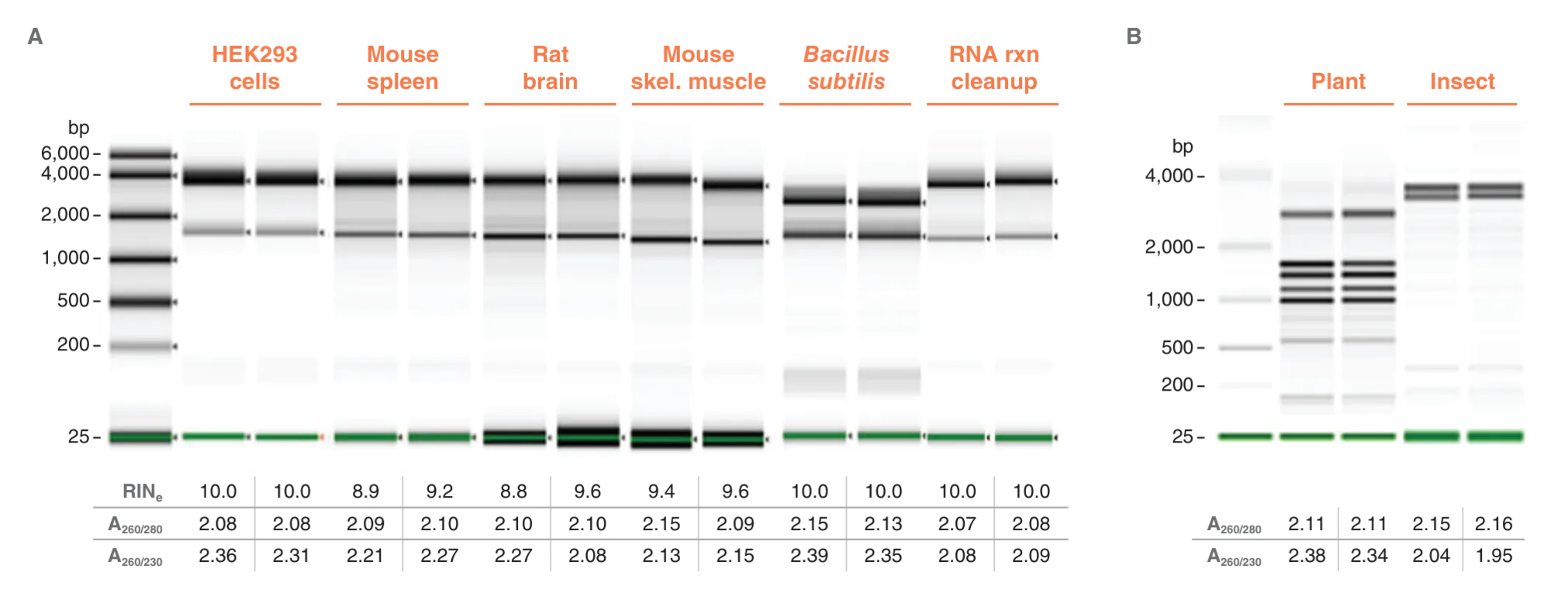
Monarch Spin RNA Isolation Kit (Mini) is a versatile solution for high-quality RNA extraction from a wide variety of sample types
RNA was extracted in duplicate using the Monarch Spin RNA Isolation Kit (Mini) from various samples representing a range of biological properties in the starting material, including fibrous, fatty and nuclease-rich mammalian tissues, tough-to-lyse samples such as Gram-positive bacteria, plants and insects, as well as RNA cleanup reactions. RNA quality was assessed using A260/A280 and A260/A230 ratios measured using a microvolume spectrophotometer (Lunatic, Unchained Labs). RNA integrity was measured using Agilent automated electrophoresis systems according to sample types. For samples with known typical RNA profiles, Agilent RNA ScreenTape was used on the TapeStation 4200 (A). For samples with known atypical RNA profiles such as plant green tissue with plastidal content, and insects with ribosomal gene breaks, Agilent Bioanalyzer 2100 used with NanoChip (B).
What our customers say about the Monarch RNA kits:
|
„Currently, a severe but small ssRNA-based plant pathogen, the citrus bark cracking viroid, is threatening hop production in Europe. Since the viroid was discovered in Germany in 2019, we are supporting the viroid monitoring and conducting degradation studies. Therefore, we needed reliable and efficient RNA extractions and compared several extraction protocols including expensive soil extraction kits. Only the Monarch Total RNA Miniprep Kit let to successful RNA extraction in all cases, thus allowing viroid detection from difficult matrices as plant residues, silage, biogas slurry, compost, and even terpenoid-rich fruit skin.“ Dr. Michael Helmut Hagemann, Researcher at the University of Hohenheim, Stuttgart Germany |
||||
 |
||||
Further information can be found in our Technical Resources section or at neb.com. Information on trademarks can be found here.