SplintR Ligase efficiently catalyzes the ligation of adjacent, single-stranded DNA splinted by a complementary RNA strand. This previously unreported activity may enable novel approaches for characterization of miRNAs and mRNAs, including SNPs. SplintR is ideally suited for many target enrichment workflows with applications in next-generation sequencing and molecular diagnostics. The robust activity of the enzyme and its affinity for RNA-splinted DNA substrates (apparent Km = 1 nM) enable sub-nanomolar detection of unique RNA species within a complex mixture, making SplintR ligase a superior choice for demanding RNA detection technologies.
Applications
- Ligation of ssDNA splinted by complementary RNA sequences
- Detection of RNA using ligation of DNA probes
- SNP or splice variant detection
- RASL-seq
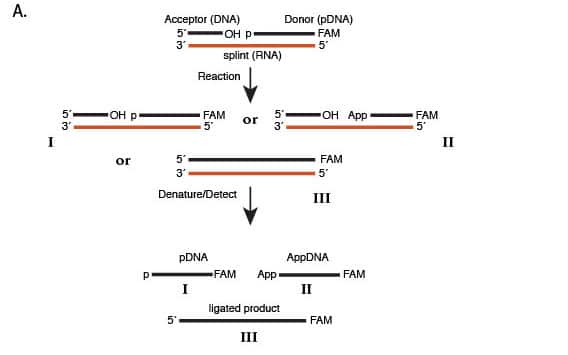
(A) Outline of the ligation assay: a 5´-phosphorylated, 3´-FAM labeled DNA “donor” oligonucleotide and an unmodified DNA “acceptor” oligonucleotide are annealed to a complementary RNA splint. This substrate is reacted with a ligase to form a mixture of unreacted starting material (I), adenylylated DNA (II), and ligated product (III). These products are denatured, separated by capillary electrophoresis and detected by fluorescence.
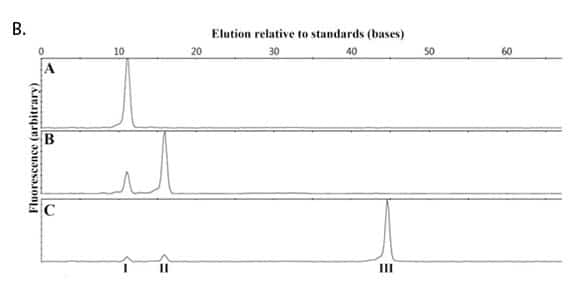
(B) Ligation of the RNA-splinted substrate in SplintR Ligase Reaction Buffer for 15 minutes at 25°C with (a) no enzyme, (b) 1 μM T4 DNA Ligase and (c) 100 nM SplintR Ligase. Indicated peaks correspond to starting pDNA (I), AppDNA (II) and ligated product (III) as determined by co-elution with synthetically prepared standards.
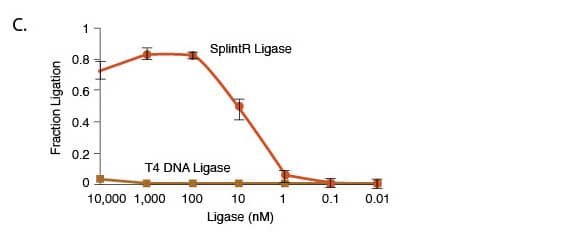
(C) The fraction of ligated product catalyzed by either SplintR Ligase or T4 DNA Ligase was analyzed by performing sets of ligations with both ligases at concentrations between 10 pM and 10 μM for 15 minutes at 25°C. SplintR Ligase is clearly much more efficient at ligation of RNA splinted DNA than T4 DNA Ligase.
Further information can be found in our Technical Resources section or at neb.com. Information on trademarks can be found here.

