Accurate quantitation of next-generation sequencing libraries is essential for maximizing data output and quality from each sequencing run.
Furthermore, for Illumina sequencing, accurate quantitation of libraries is critical to achieve optimal cluster densities, a requirement for optimal sequence output.
qPCR is considered to be the most accurate and effective method of library quantitation, providing considerably higher consistency and reproducibility of quantitation.
Amplification-based methods quantitate only those molecules that contain both adaptor sequences, thereby providing a more accurate estimate of the concentration of the library molecules that can be sequenced.
- Be confident in your quant values, as our kit provides more accurate and reproducible results than other methods and kits.
- Get up and running quickly with our easy-to-use kit, containing Library Dilution Buffer, optimized master mix and ROX dye.
- Enjoy the flexibility to use either four or six standards (included).
- Simplify your reaction setup with fewer pipetting steps and a single extension time for all libraries.
- Use with all your libraries, regardless of insert size, GC content and preparation method.
- Save money with our value pricing.
Get a free NEBNext Library Quant Kit sample!
The NEBNext Library Quant Kit delivers significant improvements to qPCR-based library quantitation for next gen sequencing.
Feature Article:
„The Quantitation Question: How does accurate
library quantitation influence sequencing?“

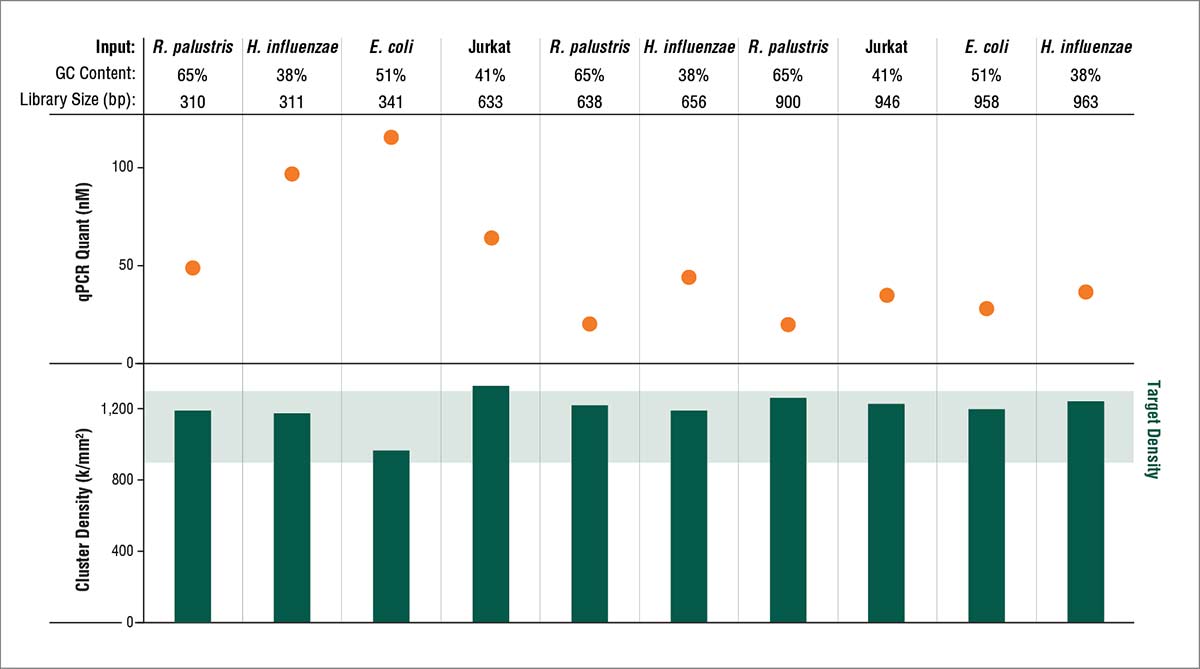
With NEBNext, optimal cluster density is achieved from quantitated libraries with a broad range of library size and GC content.
Libraries of 310–963 bp from the indicated sources were quantitated using the NEBNext Library Quant Kit, then diluted to 8 pM and loaded onto a MiSeq® (v2 chemistry; MCS v2.4.1.3). Library concentrations ranged from 7–120 nM, and resulting raw cluster density for all libraries was 965–1300 k/mm2 (ave. =1199). Optimal cluster density was achieved using concentrations determined by the NEBNext Library Quant Kit for all library sizes.
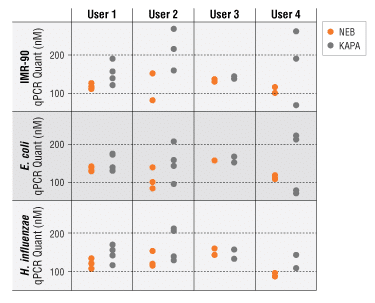
Greater reproducibility of library quantitation with the NEBNext Library Quant Kit
Three 340–400 bp libraries were quantitated by 4 different users 2–4 times using either the NEBNext or KapaTM Library Quantification Kit (Universal). A notable improvement in quantitation consistency was observed for concentrations determined by the NEBNext Kit (orange) versus those from the Kapa kit (gray).
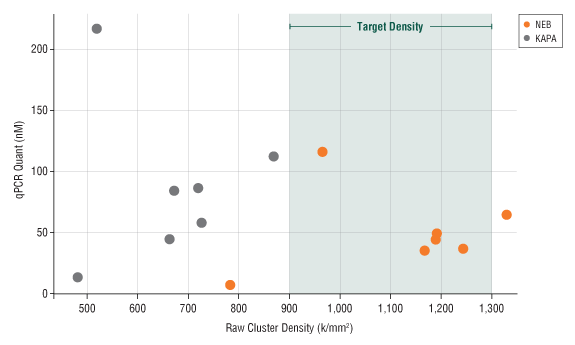
The NEBNext Library Quant Kit values enable optimal cluster densities
Seven different libraries were quantitated using either the NEBNext Library Quant Kit (orange) or the Kapa Library Quantification Kit (Universal) (gray). Undiluted library concentrations ranged from 2–200 nM. Libraries were diluted to 8 pM and loaded onto a MiSeq® instrument (v2 chemistry; MCS v2.4.1.3). Libraries quantitated with the NEBNext kit resulted in a raw cluster density average of 1160 k/mm2, directly in the optimal range of 900–1300 k/mm2. In contrast, libraries loaded based on the Kapa quantitation averaged only 660 k/mm2.
Further information can be found in our Technical Resources section or at neb.com. Information on trademarks can be found here.